Sequencing technology is playing an increasingly important role in agriculture, including crops, livestock and poultry, aquatic organisms, and more. In recent years, scientists have made great progress in the basic research of animal husbandry and molecular breeding of crops. Based on high-throughput sequencing, a series of key technologies have emerged to empower genome map construction, gene mining, and molecular breeding, thereby improving the overall competitiveness of the agricultural industry.
There are generally several types of sequencing in the agricultural sequencing field:
De novo sequencing
De novo sequencing is a method used to obtain the complete genome sequence information of the species, including different types of DNA libraries construction, sequencing, assembly and annotation with bioinformatics methods.
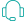
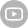
In a recent paper published by iScience titled "The chromosome-level genome for Toxicodendron vernicifluum provides crucial insights into Anacardiaceae evolution and urushiol biosynthesis", researchers from Xi'an Botanical Garden of Shaanxi Province in China successfully provided the first chromosome-level genome for Toxicodendron vernicifluum.

The lacquer tree (Toxicodendron vernicifluum) is an important tree with economic, industrial, and medicinal values. Despite its considerable significance, genetic information of the species remains scarce, which has hindered its study and utilization.
Using genomic, transcriptomic, and metabolomic techniques, researchers thoroughly screened and discovered the candidate genes responsible for urushiol and lignin production in the lacquer tree. The availability of this genomic data will be helpful in the exploration of urushiol biosynthesis, as well as molecular breeding and engineering of lacquer trees.
During the research, a total of 130.8 Gb of short reads and 106.28 Gb of ONT long reads were obtained based on DNBSEQTM and Oxford Nanopore platforms, respectively. After quality control and trimming, there were 124.4 Gb of DNBSEQTM clean reads and 105.28 Gb of ONT high-quality reads.
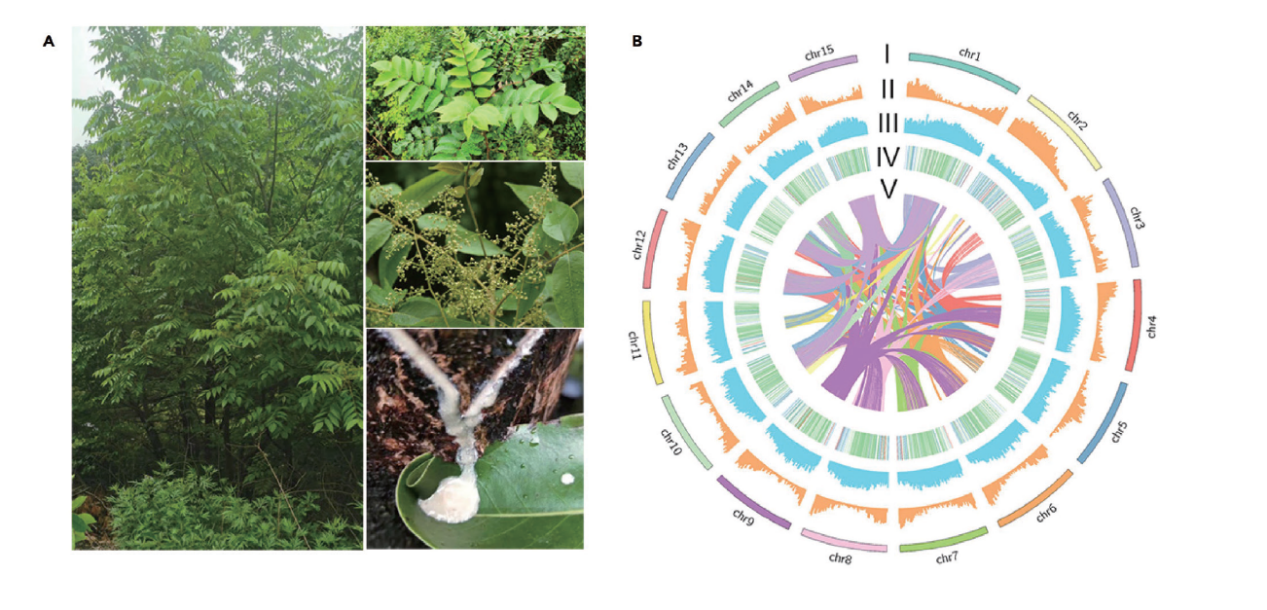
In their data, researchers found 79 orthologs containing 33 TvPKS genes involved in urushiol biosynthesis according to their co-expression profile and homological structure. Based on this, researchers drew the skeleton of the urushiol biosynthesis pathway and provided more details at the whole genome level.
DNBSEQ-G400*, the sequencing platform based on DNBSEQTM technology, helped support the assembly of the genome, including generating short reads to correct the error for long reads and producing Hi-C data for chromosome-level assembly and RNA-seq.
QTL mapping
Many agronomic traits of crops, such as quality, yield, and stress resistance, are quantitative traits. With continuous updates and improvements in molecular markers, mapping populations, and statistical analysis methods, quantitative traits locus (QTL) mapping has proved an effective approach for dissecting the genetic mechanisms of complex quantitative traits. In recent years, increasing numbers of candidate genes within QTL intervals for important agronomic traits of crops (tomato, rice, and wheat) have been isolated and cloned.
In a research published in G3: Genes, Genomes, Genetics named "Mapping of a major QTL controlling plant height using a high-density genetic map and QTL-seq methods based on whole-genome sequencing in Brassica napus", researchers from the Oil Crops Research Institute in the Chinese Academy of Agricultural Sciences set out to determine the genetic impact on plant height.

Plant height is a crucial element related to plant architecture that influences the seed yield of oilseed rape.
Traditional QTL mapping typically constructs segregating populations derived from two parental inbred lines, which is laborious, time-consuming, and costly. In recent years, a combination of QTL-seq and traditional QTL mapping has become popular for identifying QTL and involves a mutual QTL verification process in crops. In a previous study on B. napus, a major QTL underlying boron uptake was identified by QTL-seq and then molecular markers were developed based on resequencing data to fine-map the candidate gene.
Therefore, a sequencing platform with high data quality and sequencing throughput is a critical tool for agricultural genetic breeding. In this research, MGI's DNBSEQ-G400* sequencing platform was used to analyze QTL and transcriptome sequencing of specific parts of 200 rapeseed F2 isolated populations. In the end, the key QTL locus controlling plant height was identified on chromosome 10 of the rapeseed.
Whole-genome sequencing (WGS)
Whole-genome sequencing (WGS) is a comprehensive method for analyzing entire genomes. It is helpful in quickly discovering genetic variation related to important traits of animals and plants and shortening the experimental cycle of molecular breeding. WGS is now widely used for sequencing of any species, such as agriculturally important livestock, plants, or disease-related microbes.
Researchers from the National Academy of Agricultural Science in Korea shared in "Whole-genome sequences of 37 breeding line Bombyx mori strains and their phenotypes established since 1960s" the whole-genome sequences of 37 breeding line B. mori strains established over the past 60 years, along with a description of phenotypic characteristics and photos. These whole-genome sequences linked to the phenotypic characteristics of the established breeding line could be valuable resources for the understanding of B. mori genome and provide more insights into the molecular background of various phenotypes.
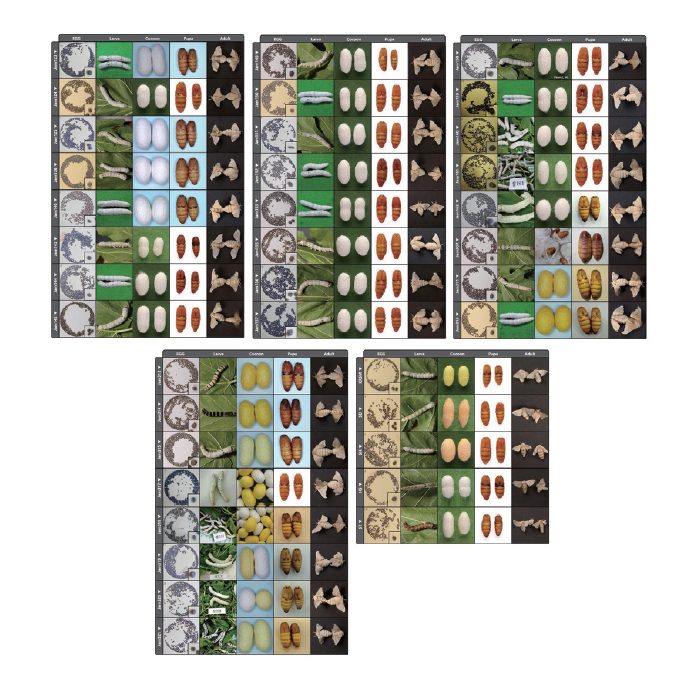


The researchers completed library construction and data generation using MGI's DNBSEQ-G400* sequencing platform and MGIEasy DNA Library Prep Kit.
Low-coverage whole genome sequencing (lcWGS)
Low-coverage whole genome sequencing (lcWGS) has emerged as a powerful and cost-effective approach for population genomic studies in both model and non-model species.
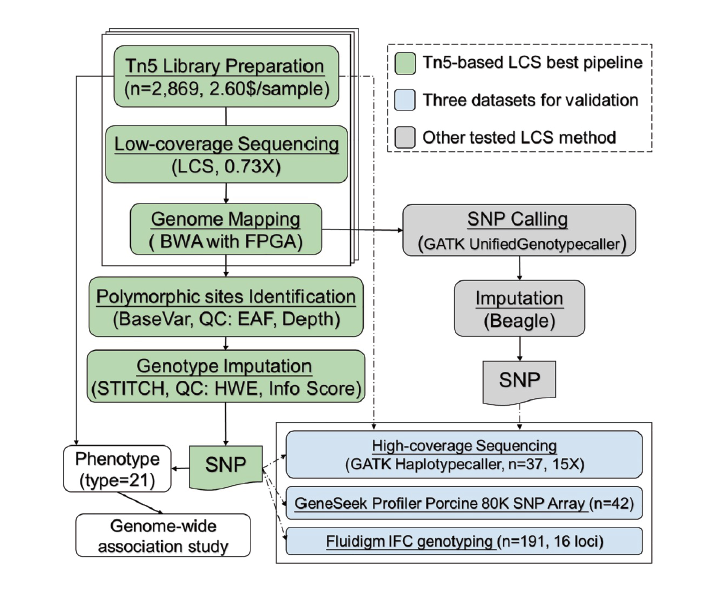
According to "Accelerated deciphering of the genetic architecture of agricultural economic traits in pigs using a low-coverage whole-genome sequencing strategy", researchers from the China Agricultural University and South China Agricultural University developed a new, highly accurate, cost- and time-efficient lcWGSmethod to obtain high-density single-nucleotide polymorphism (SNP) markers for a large Duroc pig population.
By assessing 21 important agricultural traits in commercial pig herds, they performed genome-wide association and fine-mapping analyses with high resolution and compared the results of the inheritance model in depth. Results show that artificial selection plays a significant role in altering the genetic architecture of agricultural animals, especially for loci that affect economically important traits. The lcWGS strategy had been proven useful for agricultural breeding.

The researchers completed Tn5 Library generation and sequencing on DNBSEQTM technology-based sequencing platforms DNBSEQ-G400*. DNBSEQTM significantly improves the accuracy of testing, while its low depth resequencing (Lc-WGS) genotyping is also more accurate than other methods. MGI's automatic database building scheme significantly shortens the experimental period and reduces errors commonly found in manual database building, meeting the aging requirement of molecular breeding.
From developing the economic trait markers of species and other omics research directions, to protecting the health of crops and livestock, and contributing to global agriculture, MGI provides end-to-end products for a diverse range of scenarios.
About MGI
MGI Tech Co., Ltd. (or its affiliates, "MGI"), headquartered in Shenzhen, is committed to building core tools and technology to lead life science through intelligent innovation. Based on its proprietary technology, MGI focuses on research & development, production, and sales of sequencing instruments*, reagents*, and related products to support life science research, agriculture, precision medicine and healthcare. As of December 31, 2022, MGI has more than 2,800 employees, and 36% of whom are R&D personnel. Founded in 2016, MGI operates in more than 90 countries and regions, serving more than 2,000 customers. For more information, please visit the MGI website or connect on Twitter, LinkedIn or YouTube.
*Unless otherwise informed, StandardMPS and CoolMPS sequencing reagents, and sequencers for use with such reagents are not available in Germany, Spain, UK, Sweden, Italy, Czech Republic, Switzerland and Hong Kong (CoolMPS is available in Hong Kong).
*Products are provided for Research Use Only. Not for use in diagnostic procedures (except as specifically noted)

 Sequencer Products: SEQ ALL
Sequencer Products: SEQ ALL












 Technologies
Technologies Applications
Applications Online Resources
Online Resources Data Bulletins
Data Bulletins Service & Support
Service & Support Global Programs
Global Programs Introduction
Introduction Newsroom
Newsroom Doing Business With Us
Doing Business With Us Creative Club
Creative Club













