STOmics Solutions
View the Brochure >
|
Category |
Stereo-seq Transcriptomics |
Stereo-seq Multi-omics |
||
| Product Solution |
Stereo-seq Transcriptomics |
Stereo-seq OMNI Transcriptomics |
Stereo-seq Transcriptomics for Large Chip Designs (LCD) |
|
| Detection Scope |
RNA |
RNA |
RNA |
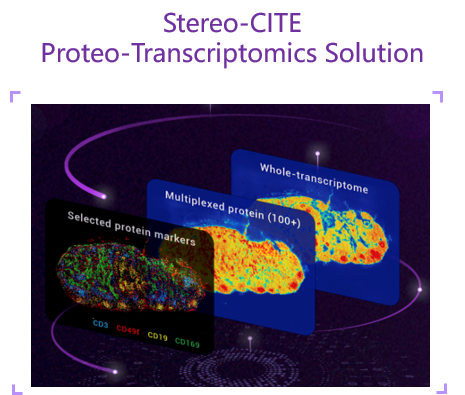
RNA + Protein |
|
Capturing Method |
Poly-A |
Random Probe |
Poly-A |
Poly-A |
| Sample Type |
Fresh Frozen (FF) |
Formalin-fixed Paraffin-embedded (FFPE) |
Fresh Frozen (FF) |
Fresh Frozen (FF) |
|
Staining Method |
ssDNA or H&E or immunofluorescence staining (mIF) |
ssDNA or H&E |
ssDNA |
DAPI |
|
Chip Size |
0.5 cm*0.5 cm, |
1 cm*1 cm |
1 cm*2 cm, 2 cm*2 cm, |
1 cm*1 cm |
| Compatible Species |
Species agnostic |
Species agnostic |
Species agnostic |
Human and mouse |
| Recommended Sequencing Amount |
0.5 cm*0.5 cm: 300-600 M reads 1 cm*1 cm: 1.5~2B reads |
3B reads |
Based on Capture Area |
Transcriptomics: 1B read Protein: 0.5-1B reads |
|
Analysis Software Version |
SAW v8.1 and StereoMap v4.1 |
SAW v8.1 and StereoMap v4.1 |
SAW v8.1 and StereoMap v4.1 |
SAW v8.1 and StereoMap v4.1 |
|
Application Highlights |
Unbiased spatial single-cell transcriptomics analysis, Combine tissue morphological information with spatial transcriptome profiling data at high resolution, RNA + low-plex protein co-detection |
True single-cell level spatial gene expression profiling on FFPE samples for diverse applications, Host-microorganism RNA CO-detection |
Unbiased spatial single-cell transcriptomics analysis with large field-of-view |
Unbiased whole transcriptome with protein multiplexing (100+ plex) simultaneously |
STOmics Platform: Your Comprehensive Solution for Empowering Multi-omics Research

- Product Files
- Technology Files
- Application Files

 Sequencer Products: SEQ ALL
Sequencer Products: SEQ ALL












 Technologies
Technologies Applications
Applications Online Resources
Online Resources Data Bulletins
Data Bulletins Service & Support
Service & Support Introduction
Introduction Newsroom
Newsroom Doing Business With Us
Doing Business With Us Creative Club
Creative Club










