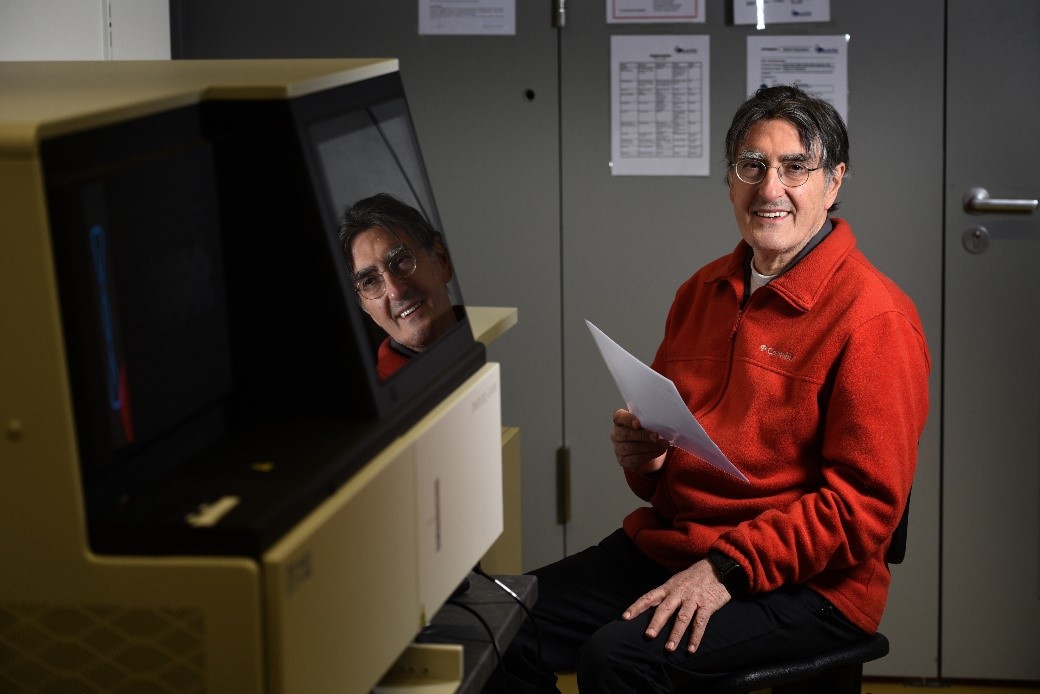
In the last four decades, Professor Dr. Hans Lehrach has been one of the greatest pioneers and leading international personalities in functional and structural genome analysis research. In particular, his scientific activities and visions have contributed to accelerating the research approaches to these disciplines, especially in Germany.
In 1978, as group leader at European Molecular Biology Laboratory in Heidelberg, Professor Dr. Lehrach contributed to develop positional cloning strategies and the first phage vectors and yeast artificial chromosomes for genomic libraries cloning. In 1987, as head of the Genome Analysis department at the Imperial Cancer Research Fund (ICRF) in London, he succeeded to map Chorea Huntington’s disease gene on chromosome 4. At ICRF, he designed and constructed the first picking and spotting array robots and implemented hybridization fingerprinting technology for genome mapping and sequencing. In 1994, he was selected to join the Max-Planck Society in Germany as Director of the Institute for Molecular Genetics in Berlin-Dahlem. During this time, he participated in the International Human Genome Sequencing Consortium finalizing the sequencing of chromosome 8 and 21. Professor Dr. Lehrach is also co-founder of several successful biotech companies in Germany.
He is currently using MGI DNBSEQ™ technology to help move the approach of clinical DNA sequencing applications from whole exome to whole genome sequencing in cancer and other pathologies. Danilo Tait, Business Development Director for MGI Europe, recently conducted an interview with Professor Dr. Lehrach.
DT: 25 years have passed since I joined your team as a PhD student at the Imperial Cancer Research Fund at Lincolns in Field, in London. Many of the researchers working there - including Rade Drmanac, MGI chief scientific officer who invented our sequencing technology - have now become key opinion leaders in genomics and are managing important research centers and core facilities. How much did the approach to genomics change since then?
HL: Quite a bit. At that point we were in the early stages of the human genome project, tooling up to sequence the first genome over 13 years, at an overall cost of more than 3 billion euros. Now the fastest machine currently available, your DNBSEQ-T7, can sequence 60 genomes in a day, at a cost of a few hundred dollars per genome. So, in a relatively short time, we have moved from knowing very little about the human genome, to being able to know much more about the differences between us as individuals than we knew about all of human biology a few decades ago.
DT: In the meantime, in 1995, you moved to Germany as a director of Max Planck Institute for Molecular Genetics in Berlin. Which differences did you encounter between the two countries, regarding the approach to genomics?
HL: The UK was (and, in a sense, still is), much
more open to genome approaches, maybe due to a perceived link between genome,
genetics and eugenics in Germany.
DT: We know today that England is the promoter of one of the most ambitious projects in the field of genomics and personalized medicine, while in Germany, people continue to discuss how to proceed with a national genome initiative. Do you think, these differences still exist between these two countries?
HL: Definitely. The UK is still way ahead. The difference in the way the healthcare system is organized (the National Health Service in the UK versus a fairly fragmented system with many different and differently organized components in Germany) might also play a role.
DT: What is the risk to Germany and the other European countries if they do not soon adopt a national genome project such as those in England, United States, France and China?
HL: The national genome projects are just one component in an overdue move to a true personalization of therapy and prevention. We are all different. We have different genomes, different environments, different disease histories, and, if we look closely enough, molecularly quite different diseases. It is therefore not surprising that we react very differently to therapies or preventive measures, with sometimes disastrous consequences for us as individuals and as societies. This contributes, among other factors, to the close to 200,000 deaths from adverse drug reactions every year in Europe alone, as well as to the enormously high (and still increasing) healthcare costs in many regions (4.5 billion euros per day in Europe. On one hand, many infected (and infectious) individuals show no or only very subtle symptoms, making it very hard to control the pandemic. On the other hand, a significant fraction of the infected suffer disastrous consequences, requiring intensive medical care, leaving long-term medical consequences, and, in some cases, leading to death, making it absolutely essential, that we do so. A better understanding why individuals react so differently to the same infection could provide much better tools to stop the infection process, and to avoid the serious medical consequences in a fraction of the infected individuals
DT: We come to the Covid-19 crisis now. Strangely, if we look at the number of people who have died during the crisis, in Germany there has been a better response to the pandemic than in England. What is the reason, in your opinion?
HL: It is particularly astonishing, since, in principle, we would expect a centralized system like the NHS to be able to react much faster to such a challenge, than the much more diversified system in Germany, but, unfortunately, centralized systems are also much more sensitive to suboptimal decisions made at the top, more out of public relations considerations, than a reality-driven decision process. A former quantum-physicist as head of the government probably helps a lot in this respect.
DT: Recently, in several articles published in the German press, you and George Church as co-author, have proposed the surveillance approach by sequencing the virus and the host as the solution to the pandemic, while we are waiting for the vaccine. Could you please explain the idea behind this approach?
HL: In contrast to HIV, for example, SARS-CoV-2 does not seem to be able to survive in individuals for longer periods, since the infected persons either get cured or, in the worst case, die. This ‘Achilles’ heel’ of the virus could, in principle, be used to eliminate it again regionally, nationally or even across most (or all) of the world by carrying out a few cycles of synchronized, population-wide tests, and putting all infected (and those who refuse to be tested) in quarantine, till they are not infectious any more. This would, for a short period, bring the R factor, the number of individuals an infected person infects, to zero, and essentially eliminate the virus from the area. If this is, for example, carried out in the Schengen Area, combined with stringent testing/short quarantine at the external borders, we all could (again) rely on the fact that the people we interact with in our private life or our job are not infected, and live and travel freely within the areas, where the virus has been eliminated.
Testing all Europeans, for example, once a week for five weeks is obviously a non-trivial logistics problem, but just like the landing on the moon, not beyond human ingenuity. It is also much less costly than extending the current situation until we get a vaccine, which might take a long time, or might never happen at all (we are still waiting for the vaccine against HIV promised to us in the ‘80s of the last century).
At Alacris Theranostics (www.alacris.de), a spinoff of my former department at the Max Planck Institute for Molecular Genetics, we are, for example, working on a fully scalable testing strategy), which might, in theory (dependent on access to enough test tubes) be able to carry out population-wide tests nationally or supra-nationally, based on a highly scalable concept of sample processing, combined with NGS techniques to generate tens of billions of reads in a day.
DT: Do you think laboratories are ready for this package and have enough tools and financial support to do it? Sequencing is still considered an expensive screening approach.
HL: The pure sequencing part of the test would probably be in the cent range, with overall costs probably in the range of a few euros, an order of magnitude or more below the cost of the tests used currently. If we can end this nightmare at a cost of less than the salary for one hour of work for every European, rather than having them all out of the economic process for month, the costs have to be far, far cheaper than continuing the current compromise between too many deaths and too much economic damage.
DT: We have also seen how viral infection has affected, differently, various age groups and people at risk. Is this further proof that the personalized study of medicine, in this case related to the pathologies generated by the virus, is the right way to go?
HL: Truly personalized medicine will be inherently much more powerful and much more cost effective than the current situation, moving slowly away from the (up to a short time ago, unavoidable) one-size-fits-all strategy, which still dominates large areas of medicine. As I mentioned above, this is particularly the case for COVID-19, in which the enormous differences in response to infection play a major role in the difficulties to handle this unprecedented pandemic.
DT: Last year MGI Tech, the technology branch of the MGI, announced the launch of a new sequencer - the DNBSEQ-T7 - which can sequence 60 human genomes in 24 hours at very low sequencing costs. Which impact can this technology and its cost have on the use of genomics in personalized medicine?
HL: The T7 is a fantastic machine, twice as fast, significantly cheaper to run and with a number of other performance advantages than the closest competitor. We count on this type of progress in machine capabilities in our plans for the next steps in therapy personalization, initially in oncology, but then extending to more and more other areas or therapy and prevention, as well as our plans to get COVID-19 under control.
DT: You are currently using MGI sequencing technology at Alacris Theranostics, the company you founded in Berlin. Can you briefly explain which role DNA sequencing plays in the innovative “Digital Twins technology” at Alacris and why did you decide to rely on MGI technology for this?
HL: To identify the individually optimal therapy for every patient, we have to follow the example of almost all other areas of our existence: in complex situations, we cannot avoid making mistakes, but we can make them safely, quickly and cheaply on computer models of the real situation. For this, we have to carry out deep molecular analyses of the patient, the diseased tissue and other aspects of the patient likely to affect the disease and the therapy response (e.g. the individual immune system), information also needed for the precision medicine approach to therapy choice. To be able to convert this wealth of diverse data into predictions, how the individual patient will respond to a specific therapy, we try to develop Digital Twins, expected to correspond closely to the individual patient, to be able to identify their likely effects and side effects, without endangering the real patient. All this is only possible, using high performance, cost-effective sequencing machines, like the MGI instrument we use heavily in these analyses.
DT: And finally, how do you imagine the world of genomics and personalized medicine in 25 years, when will we meet again to take stock of the situation?
HL: In 25 years nobody will understand anymore, why it was not obvious from the beginning that medicine had to be truly personalized, based on extensive characterization and modelling of our biology. Nobody will understand that we did not have individual Digital Twins, guiding therapy and prevention, but also taking our individual biology into account in sport and wellness. Nobody will understand that the development of new drugs not using Digital Twin type models in development and approval, could have been a process typically taking much more than a decade, and costing billions. We will have moved from the Ptolomean view (everybody is similar, give or take a few epicycles) to a sort of Copernican view: we are all different, and able to make the right decisions in spite of this, and will have forgotten, the ‘dark ages’ we are living in now, just as much, as we have forgotten the ‘dark ages’ of our existence before the internet.

 Sequencer Products: SEQ ALL
Sequencer Products: SEQ ALL











 Technologies
Technologies Applications
Applications Online Resources
Online Resources Data Bulletins
Data Bulletins Service & Support
Service & Support Introduction
Introduction Newsroom
Newsroom Doing Business With Us
Doing Business With Us Creative Club
Creative Club













