MGISEQ-200 Used to Confirm Suspected Cases of COVID-19
"The MGISEQ-200 DNBSEQ platform helped us to fight against COVID-19 in terms of ultra-high sensitivity detection of complex infections, virus mutation monitoring, tracing, pathogenesis, prediction, and assistance in clinical treatment."
——Bamrasnaradura Infectious Diseases Institute, Department of Disease Control, Thailand
In Thailand the rapid and effective development of epidemic prevention and control work has helped to keep the cumulative number of confirmed cases of COVID-19 with just over 3,000. The local disease control system uses a combination of RT-PCR nucleic acid detection technology and high-throughput sequencing technology to confirm the diagnosis of suspected COVID-19 cases and also to launch follow-up research and analysis. As part of this approach, the high-throughput sequencing program adopted by the Bamrasnaradura Infectious Diseases Institute, Department of Disease Control is based on the MGISEQ-200 platform and its rapid identification system for pathogens, known as PFI (Pathogen Fast Identification).
In the early stages of the epidemic, BIDI mainly used RT-PCR for nucleic acid detection of COVID-19, but some suspected samples COVID-19 fell into a gray area based on the Ct value of the RT-PCR test results, corresponding to the level of gene expressed. In order to improve the ability of diagnosis, the BIDI introduced MGISEQ-200 and its matching PFI (Pathogen Fast Identification) system. Under the field training (Figure 2) and guidance of technical engineers from MGI, the BIDI quickly launched high-throughput sequencing technology for virus detection.
Deep sequencing with the MGISEQ-200 was used to confirm the suspected samples with a RT-PCR Ct value that was difficult to give a clear diagnosis. One sample with RT-qPCR Ct value of 39.47 was confirmed as a positive case using this method, demonstrating the advantages of massively parallel sequencing. Currently, several positive cases have been confirmed and many more such samples are being tested right now.
Using high-depth metagenomic sequencing based on MGISEQ-200 has enabled the BIDI to achieve rapid and accurate diagnosis. Among the samples submitted, various extreme samples will appear, including samples with a Ct value of nearly 40, which is close to the threshold of RT-PCR detection; and samples that are negative for the first RT-PCR test and positive for the second. To further confirm the diagnosis, the BIDI used MGISEQ-200 to perform high-depth metagenomic sequencing on suspected samples, resulting in a total of 429.1M reads with a Q30 of 90% (Figure 1). The data from the machine can be used directly to automatically enable the supporting MGI pathogen rapid identification system for analysis. The analysis results indicate that the number of COVID-19 reads detected in 2 suspected samples is 21 and 9. These results show that for samples with low viral load or trace viral load, metagenomic sequencing has certain advantages, which can improve sensitivity, reduce false negatives, and serve as an important complement to RT-PCR testing.
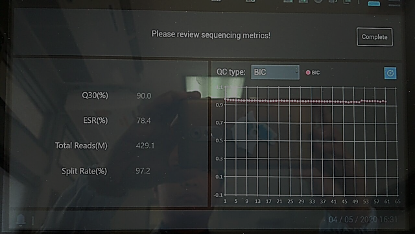
Figure 1 - MPS result of suspected samples. Photo by BIDI.

Figure 2 – MGI field application scientists have provided training to BIDI.
Our local partner in Thailand for sales and support of the MGISEQ-200 packages is: PCL holding group ; 601, 603 Soi Charatlarp, Sirinthorn Road Bangplad Bangkok 10700
Tel. +66 2096 2002 Email. info@pclholding.com

 Sequencer Products: SEQ ALL
Sequencer Products: SEQ ALL











 Technologies
Technologies Applications
Applications Online Resources
Online Resources Data Bulletins
Data Bulletins Service & Support
Service & Support Introduction
Introduction Newsroom
Newsroom Doing Business With Us
Doing Business With Us Creative Club
Creative Club